Manganese »
PDB 2ayl-2ce4 »
2bvl »
Manganese in PDB 2bvl: Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion
Protein crystallography data
The structure of Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion, PDB code: 2bvl
was solved by
D.J.Reinert,
T.Jank,
K.Aktories,
G.E.Schulz,
with X-Ray Crystallography technique. A brief refinement statistics is given in the table below:
| Resolution Low / High (Å) | 30.00 / 2.2 |
| Space group | P 21 21 21 |
| Cell size a, b, c (Å), α, β, γ (°) | 63.010, 95.797, 112.915, 90.00, 90.00, 90.00 |
| R / Rfree (%) | 19.7 / 24.4 |
Other elements in 2bvl:
The structure of Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion also contains other interesting chemical elements:
| Bromine | (Br) | 48 atoms |
| Tantalum | (Ta) | 24 atoms |
Manganese Binding Sites:
The binding sites of Manganese atom in the Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion
(pdb code 2bvl). This binding sites where shown within
5.0 Angstroms radius around Manganese atom.
In total only one binding site of Manganese was determined in the Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion, PDB code: 2bvl:
In total only one binding site of Manganese was determined in the Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion, PDB code: 2bvl:
Manganese binding site 1 out of 1 in 2bvl
Go back to
Manganese binding site 1 out
of 1 in the Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion
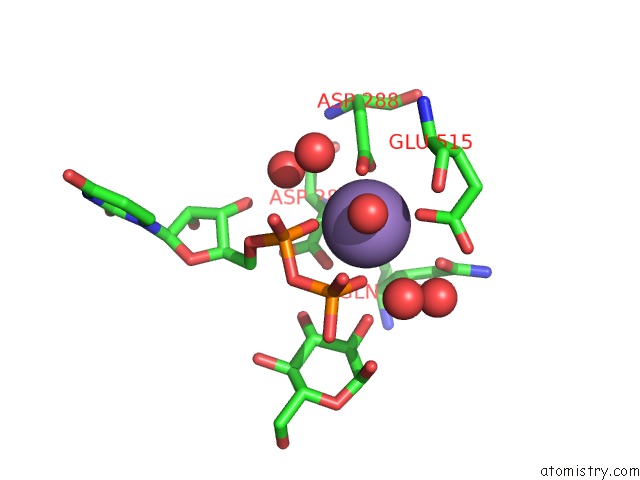
Mono view
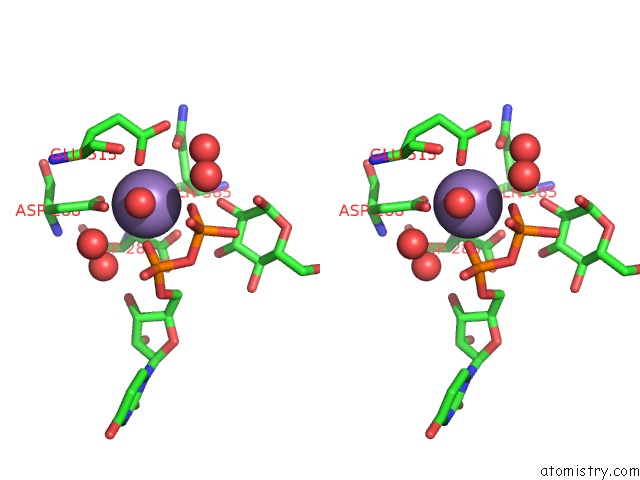
Stereo pair view

Mono view

Stereo pair view
A full contact list of Manganese with other atoms in the Mn binding
site number 1 of Crystal Structure of the Catalytic Domain of Toxin B From Clostridium Difficile in Complex with Udp, Glc and Manganese Ion within 5.0Å range:
|
Reference:
D.J.Reinert,
T.Jank,
K.Aktories,
G.E.Schulz.
Structural Basis For the Function of Clostridium Difficile Toxin B. J.Mol.Biol. V. 351 973 2005.
ISSN: ISSN 0022-2836
PubMed: 16054646
DOI: 10.1016/J.JMB.2005.06.071
Page generated: Sat Aug 16 09:50:30 2025
ISSN: ISSN 0022-2836
PubMed: 16054646
DOI: 10.1016/J.JMB.2005.06.071
Last articles
Mo in 5O5WMo in 6GB4
Mo in 6GBC
Mo in 6FW2
Mo in 6ETF
Mo in 6GAX
Mo in 6CZA
Mo in 6CZ9
Mo in 6CZ8
Mo in 6CZ7